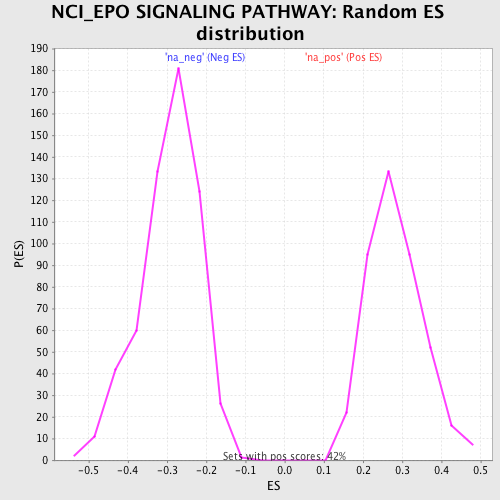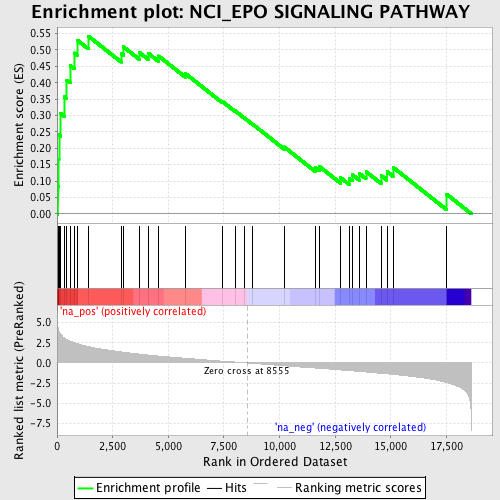
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | NCI_EPO SIGNALING PATHWAY |
| Enrichment Score (ES) | 0.5422996 |
| Normalized Enrichment Score (NES) | 1.9275196 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.034292214 |
| FWER p-Value | 0.336 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BCL2 | 39 | 4.461 | 0.0846 | Yes | ||
| 2 | RAPGEF1 | 50 | 4.258 | 0.1668 | Yes | ||
| 3 | SH2B3 | 85 | 3.913 | 0.2409 | Yes | ||
| 4 | BTK | 166 | 3.564 | 0.3059 | Yes | ||
| 5 | PTPN6 | 332 | 3.080 | 0.3568 | Yes | ||
| 6 | MAPK14 | 438 | 2.913 | 0.4078 | Yes | ||
| 7 | JAK2 | 591 | 2.705 | 0.4521 | Yes | ||
| 8 | PLCG2 | 775 | 2.497 | 0.4908 | Yes | ||
| 9 | GRB2 | 924 | 2.371 | 0.5289 | Yes | ||
| 10 | EPO | 1405 | 2.019 | 0.5423 | Yes | ||
| 11 | TEC | 2878 | 1.359 | 0.4895 | No | ||
| 12 | VAV2 | 2980 | 1.326 | 0.5098 | No | ||
| 13 | PLCG1 | 3717 | 1.086 | 0.4913 | No | ||
| 14 | CBL | 4089 | 0.980 | 0.4904 | No | ||
| 15 | STAT5A | 4560 | 0.854 | 0.4817 | No | ||
| 16 | SOS1 | 5763 | 0.579 | 0.4283 | No | ||
| 17 | PIK3R1 | 7425 | 0.214 | 0.3431 | No | ||
| 18 | LYN | 7998 | 0.109 | 0.3144 | No | ||
| 19 | NFKB1 | 8403 | 0.033 | 0.2933 | No | ||
| 20 | EPOR | 8789 | -0.047 | 0.2735 | No | ||
| 21 | GAB1 | 10210 | -0.353 | 0.2040 | No | ||
| 22 | SOCS3 | 11623 | -0.637 | 0.1404 | No | ||
| 23 | STAT1 | 11783 | -0.667 | 0.1448 | No | ||
| 24 | RAP1A | 12745 | -0.870 | 0.1100 | No | ||
| 25 | HRAS | 13137 | -0.949 | 0.1074 | No | ||
| 26 | STAT5B | 13272 | -0.980 | 0.1192 | No | ||
| 27 | BCL2L1 | 13603 | -1.056 | 0.1220 | No | ||
| 28 | TRPC6 | 13886 | -1.121 | 0.1286 | No | ||
| 29 | MAPK8 | 14563 | -1.284 | 0.1171 | No | ||
| 30 | CRKL | 14828 | -1.344 | 0.1290 | No | ||
| 31 | SHC1 | 15101 | -1.412 | 0.1418 | No | ||
| 32 | PTPN11 | 17514 | -2.432 | 0.0593 | No |